|
The area of the promoters is of great interest as researchers pay great attention to their importance not only in developmental gene expression but also in environmental response. It is essential to identify the promoter regions in a genome, as it can aid in illuminating the genome’s regulatory mechanism and disease-causing variants within cis-regulatory elements. Sigma70 promoter comprises two well-defined short sequences located at-10 and-35 base pairs upstream of TSS, known as pribnow box and-35 region, respectively ( Paget and Helmann, 2003). Sigma70 factor is a crucial factor as it regulates the transcription of most of the housekeeping genes and responsible for the most of the DNA regulatory functions. The number associated with each sigma factor represents the molecular weight. There are various types of sigma factors responsible for different functions, such as sigma54 controls the transcription of genes responsible for the modulation of cellular nitrogen levels, sigma38 regulates the stationary phase genes, sigma32 regulates heat-shock genes, and sigma24 and sigma18 controls the extra-cytoplasmic functions ( Paget, 2015). In prokaryotes, promoters are recognized by the holoenzyme, which is made up of RNA polymerase and a related sigma factor. Promoters are generally located at the upstream of genes’ transcription start sites (TSS) responsible for the switching on or off the respective gene. Promoters and enhancers regulate the fate of a cell by regulating the expression of the genes. We have developed a method, Sigma70Pred, which is available as webserver and standalone packages at. Our model successfully predicted constitutive promoters with accuracy of 81.46% on an independent dataset. In order to check the robustness of the model, we have tested our model on the independent dataset made by using RegulonDB10.8, which included 1,134 sigma70 and 638 non-promoters, and able to achieve accuracy of 90.41% with AUROC of 0.95. Our SVM based model achieved maximum accuracy 97.38% with AUROC 0.99 on training dataset, using 200 most relevant features. We have generated a wide range of features around 8,000, which includes Dinucleotide Auto-Correlation, Dinucleotide Cross-Correlation, Dinucleotide Auto Cross-Correlation, Moran Auto-Correlation, Normalized Moreau-Broto Auto-Correlation, Parallel Correlation Pseudo Tri-Nucleotide Composition, etc. 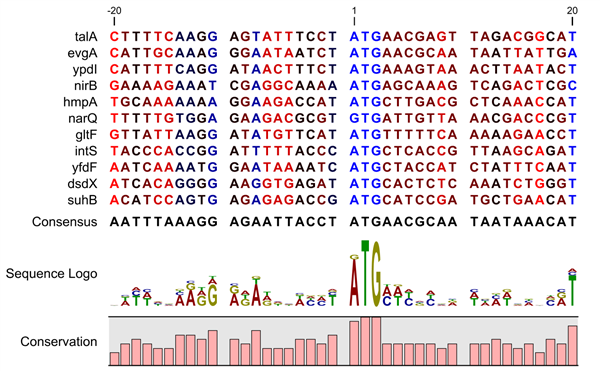 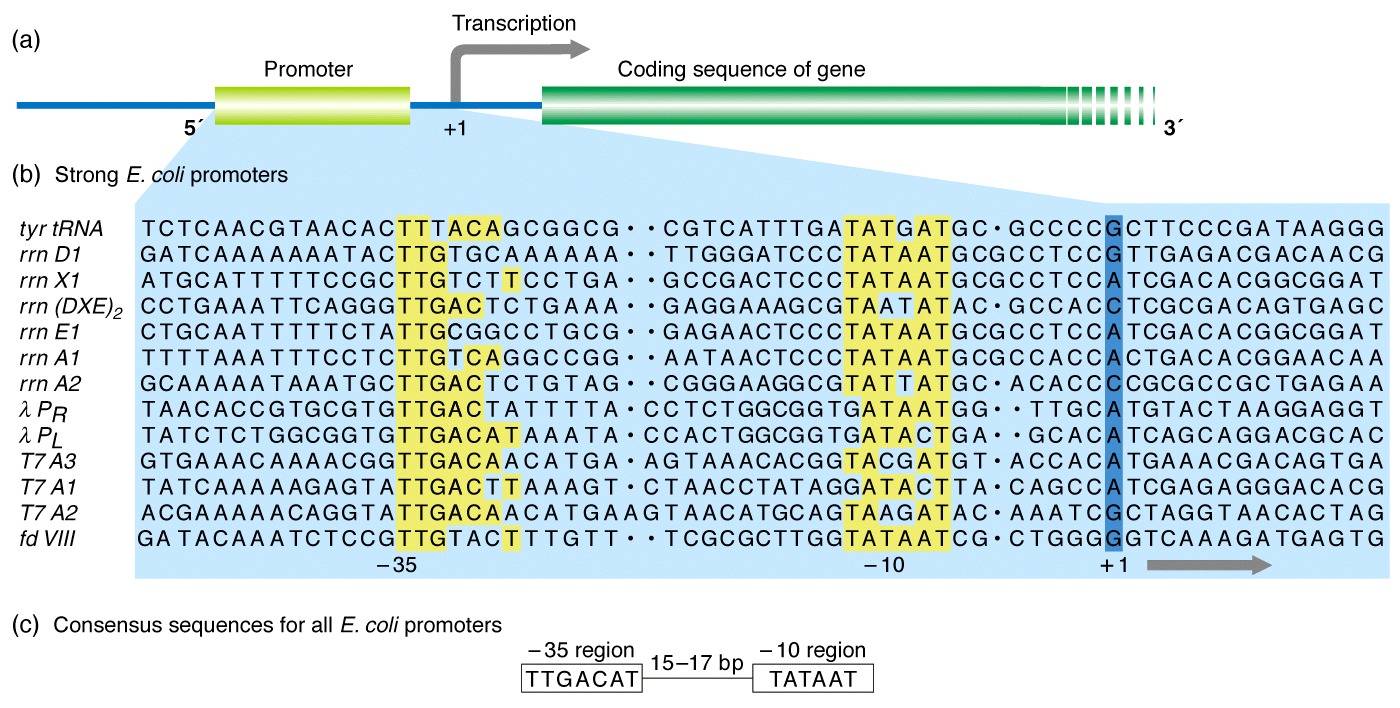
In this study, we trained and evaluate our models on a dataset consists of 741 sigma70 promoters and 1,400 non-promoters. One of the major challenges is to predict the sigma70 promoter or sigma70 factor binding site with high precision. Sigma70 factor plays a crucial role in prokaryotes and regulates the transcription of most of the housekeeping genes.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
